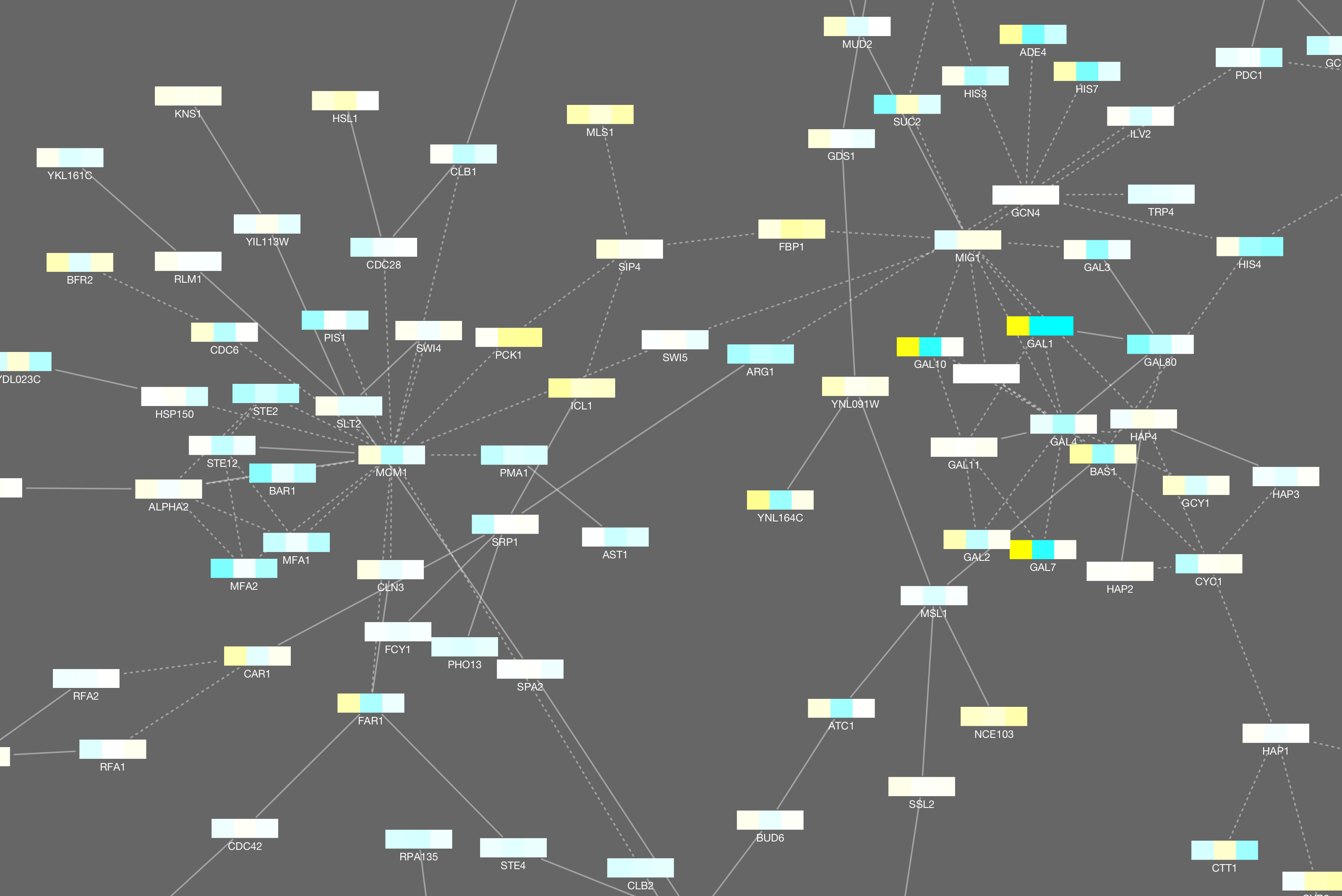
The initial set is designed to work with public bioinformatics SPARQL endpoints provided by the EBI and Bio2RDF. When you first install the app, it comes pre-configured with a basic set of SPARQL queries, although it’s possible to provide your own set. When you click on them, a query is sent off in the background, and the result is mapped to your network according to certain rules. Each item in the search menu, and each item in the context (right-click) menu, is backed by one SPARQL query. In the background, the General SPARQL app maintains a list of SPARQL queries. Watch this screencast and it will start to make sense: And you can continue this process, jumping from one entity to the next. Or the gene that encodes for your protein. Or all the reactions that your protein is part of. For example, all the pathways that are related to your protein. You can then right-click on this node to pull in related entities. The app places a single node in your network. Edges can represent any type of relation between those entities.įor example, you can start by searching for a protein of interest. Nodes can represent various biological entities, such as: a pathway, a protein, a reaction, or a compound. Nodes and edges can be added to a network in a piecemeal fashion. The app lets you build a network step by step. The General SPARQL app is one of the new ways to present triple data. We shouldn’t expect scientists to write SPARQL queries anymore than we expect them to carry adjustable pliers to a restroom visit. The problem is that SPARQL is supposed to be only a piece of plumbing at the bottom of a software stack. The problem isn’t that SPARQL is hard to use per se (it’s really rather plain and sensible). Integrating data is only half of the problem, we also have to present that data. Triple stores are great for data integration, but you still have to figure out how to put that data in the hands of scientists. But how do we put that data in the hands of scientists?Īt General Bioinformatics we put data in triple stores, and use SPARQL to query that data. In this case, the only you can do is to ask for assistance of a professional staff.We can now easily solve the problem of bioinformatics data integration. If the problem with the CYS file has not been solved, it may be due to the fact that in this case there is also another rare problem with the CYS file. If you are sure that all of these reasons do not exist in your case (or have already been eliminated), the CYS file should operate with your programs without any problem. Drivers of equipment used by the computer to open a CYS file are out of date.The computer does not have enough hardware resources to cope with the opening of the CYS file.The CYS file which is being opened is infected with an undesirable malware.Incomplete installation of an application that supports the CYS format. 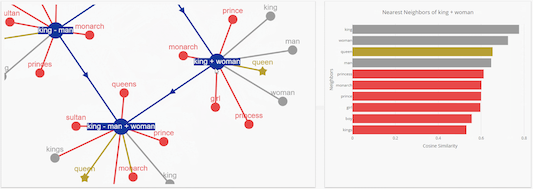
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
